Domain-Specific Pretrained Encoder Transformers for the Identification of Methodologically Rigorous Systematic Reviews: A Retrospective Modeling Study
Abstract
Fangwen Zhou,1 Muhammad Afzal,2 Rick Parrish,1 Ashirbani Saha,3 Wael Abdelkader,1 R. Brian Haynes,1 Alfonso Iorio,1,4 Cynthia Lokker1
Objective
Systematic reviews are considered one of the strongest levels of evidence, providing essential information for clinical guidelines and bedside practice. However, the broad spectrum of review articles, including narrative reviews and other types, complicates the identification of high-quality systematic reviews.1 This study aims to fine-tune and evaluate pretrained encoder transformers to identify high-quality systematic reviews from review articles using a large, reputable dataset.
Design
Review articles from McMaster’s Premium Literature Service (PLUS) were retrieved.2 Articles were considered rigorous if they were systematic reviews that (1) stated the clinical topic; (2) described methods, including databases searched and inclusion criteria; (3) searched ≥1 major database; (4) reported the number of retrieved, reviewed, and included articles; and (5) did not exclude randomized clinical trials for treatment, primary prevention, quality improvement, or economics reviews; or included “inception cohort” for prognosis reviews.3 Ground truth was established by research methodology experts using article full texts. Articles from 2003 to 2023 were randomly split 80:10:10 into training, validation, and testing sets. Articles in 2024 were used for external testing. A grid search of 7 pretrained models, 3 learning rates, 5 batch sizes, and 3 random seeds, with or without class weight adjustments, was conducted. Models were trained for ≤10 epochs, with an early stopping patience of 3. Titles and abstracts were used as inputs. Those longer than 512 tokens were truncated, and those shorter were padded. The model that achieved the lowest log loss on the validation set was further evaluated. A threshold of ≥0.50 was used for classification. Bootstrapping of 1000 iterations was used to estimate 95% confidence intervals.
Results
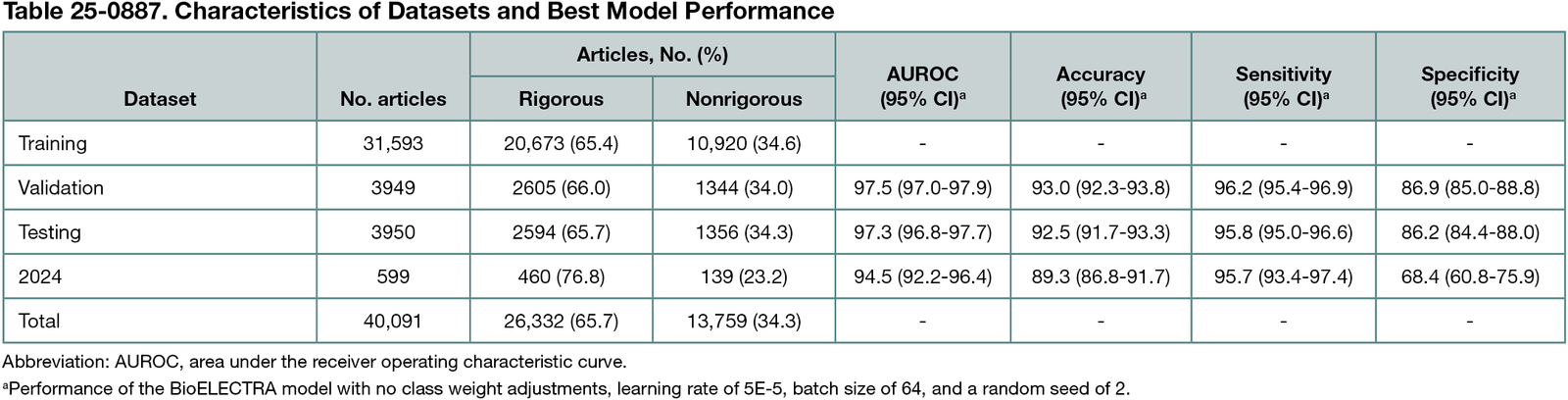
Among 40,091 reviews, 26,332 (65.7%) were considered rigorous and 31,593 were used for training. Of the 630 fine-tuned models, BioELECTRA with no class weight adjustments, learning rate 5E-5, batch size 64, and seed 2 had the lowest validation log loss of 0.1840. Table 25-0887 details the characteristics of the datasets and the model’s performance, achieving >94% area under the receiver operating characteristic curve and >95% sensitivity. In review articles published in 2024, there was a notable drop in specificity and mild degradation in other metrics.
Conclusions
Pretrained encoder transformer models, such as BioELECTRA, demonstrate robust performance in identifying rigorous systematic reviews based on PLUS’s criteria. These models can streamline the identification of high-quality evidence, reducing manual effort and enhancing the efficiency of evidence curation. Performance degradation in articles in 2024 may be due to changes in article structure and content over time. Additionally, generalizability to other datasets is unknown. Future efforts should establish standardized benchmarking datasets for systematic reviews, develop models for tools, such as AMSTAR 2, and account for temporal data drift to better support knowledge translation.
References
1. Shaheen N, Shaheen A, Ramadan A, et al. Appraising systematic reviews: a comprehensive guide to ensuring validity and reliability. Front Res Metr Anal. Published online December 21, 2023. doi:10.3389/frma.2023.1268045
2. Holland J, Haynes RB; McMaster PLUS Team Health Information Research Unit. McMaster Premium Literature Service (PLUS): an evidence-based medicine information service delivered on the Web. AMIA Annu Symp Proc. 2005;2005:340-344. https://www.ncbi.nlm.nih.gov/pubmed/16779058
3. Methodological criteria. McMaster University Health Information Research Unit. Accessed August 19, 2024. https://hiruweb.mcmaster.ca/hkr/what-we-do/methodologic-criteria
1Health Information Research Unit, Department of Health Research Methods, Evidence, and Impact, Faculty of Health Sciences, McMaster University, Hamilton, Ontario, Canada, lokkerc@mcmaster.ca; 2Department of Computing, Faculty of Computing, Engineering and the Built Environment, Birmingham City University, Birmingham, UK; 3Department of Oncology, Faculty of Health Sciences, McMaster University, Hamilton, Ontario, Canada; 4Department of Medicine, Faculty of Health Sciences, McMaster University, Hamilton, Ontario, Canada.
Conflict of Interest Disclosures
McMaster University, a nonprofit public academic institution, has contracts through the Health Information Research Unit under the supervision of Alfonso Iorio and R. Brian Haynes. These contracts involve professional and commercial publishers to provide newly published studies, which are critically appraised for research methodology and assessed for clinical relevance as part of the McMaster Premium Literature Service (McMaster PLUS). Cynthia Lokker and Rick Parrish receive partial compensation, and R. Brian Haynes is remunerated for supervisory responsibilities and royalties. Ashirbani Saha, Fangwen Zhou, Muhammad Afzal, and Wael Abdelkader are not affiliated with McMaster PLUS.
Funding/Support
Fangwen Zhou was funded through the Mitacs Business Strategy Internship grant (IT42947) with matching funds from EBSCO Canada.
Role of the Funder/Sponsor
The funders were not involved in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the abstract; and decision to submit the abstract for presentation.
Acknowledgment
We thank the Digital Research Alliance of Canada for its computational resources.
