Detection of Open Science Practices in Major Medical Journals: A Survey and Diagnostic Accuracy of Automatic Tools Using Sensitivity and Specificity
Abstract
Constant Vinatier,1 Ayu Putu Madri Dewi,2 Gwénaël Dumont,1 Tracey Weissgerber,3 Vladislav Nachev,3 Gowri Gopalakrishna,2,3,4 Maud Scheidecker,1 François-Joseph Arnault,1 Nicholas J. DeVito,5 Guillaume Freyermuth,6 Mathieu Acher,6,10 Gauthier Le Bartz Lyan,6 Inge Stegeman,7,8 Mariska M. G. Leeflang,2 F. Naudet9,10
Objective
Despite open science policies in major biomedical journals, adherence remains uncertain. This study evaluated automated tools, from regular expressions to large language models (LLMs), for assessing core open science practices in leading biomedical journals.
Design
We retrospectively assessed research articles from a sample of 10 major generalist medical journals (Annals of-Internal Medicine, BMJ, BMC Medicine, Canadian Medical Association Journal [CMAJ], JAMA, JAMA Network Open, Lancet, Nature Medicine, New England Journal of Medicine, and PLoS Medicine) from 2020 to 2023. Articles were retrieved via PubMed using a Peer Review of Electronic Search Strategies (PRESS) search strategy. The database comprised random samples of 103 randomized controlled trials (RCTs), 98 meta-analyses (MAs), and 111 other research articles (RAs). We evaluated 13 open science practices, including study registration, data sharing, and protocol sharing (open access or upon request). Each article was evaluated by 2 independent raters, with any disagreements resolved by a third rater. Seven different automated tools—rtransparent, oddpub, ctRegistries, ContriBot, DataSeer, SciScore, and an LLM (Llama 3-70B)—were used. Diagnostic accuracies were estimated using sensitivities, specificities, F1 scores, and LR+ and LR-.
Results
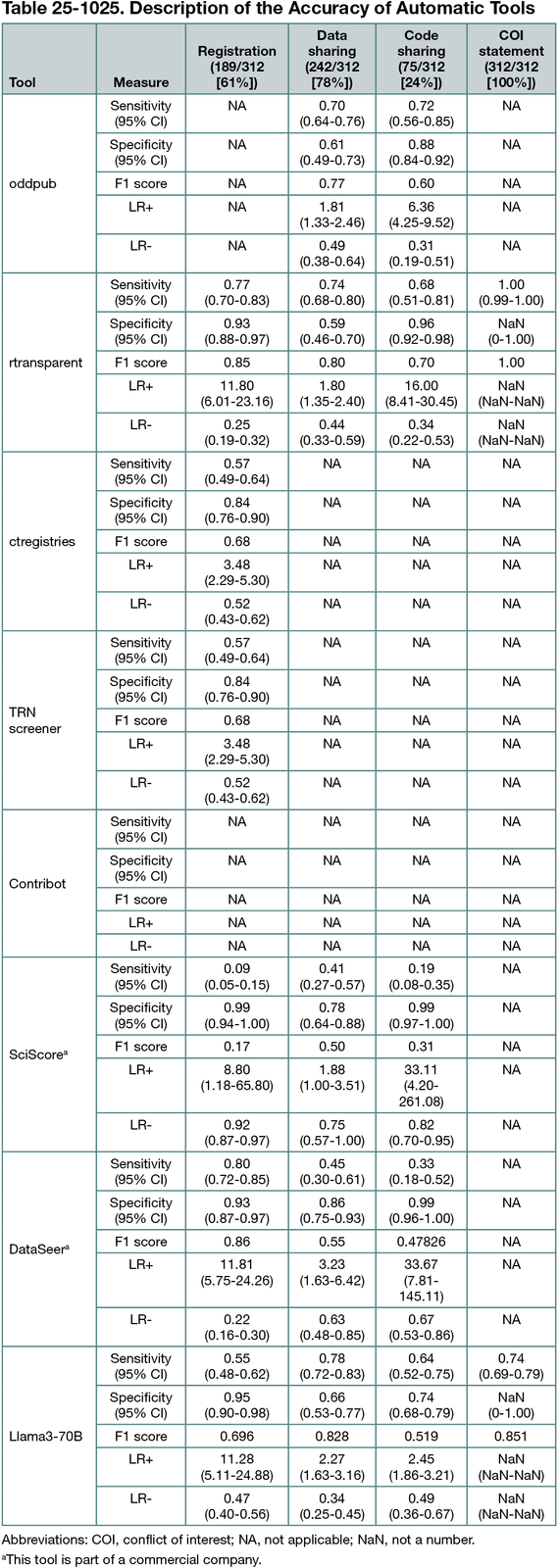
Manual extraction in the 312 articles identified registration in 98% (101/103) of RCTs, 69% (68/98) of MAs, and 18% (20/111) of RAs. Open data were present in 6% (6/103) of RCTs, 36% (35/98) of MAs, and 13% (15/111) of RAs and accessible upon request in 78% (80/103), 41% (40/98), and 59% (66/111), respectively. Protocols were openly available in 84% (87/103) of RCTs, 64% (63/98) of MAs, and 20% (22/111) of RAs and accessible upon request in 5% (5/103), 3% (3/98), and 3% (3/111), respectively. The accuracy of automated tools varied depending on the practice evaluated, with F1 scores ranging from 1.00 (Conflict of Interest statement, rtransparent) to 0.16 (SciScore, registration). For study registration, a simple tool using regular expressions, such as rtransparent, demonstrated good sensitivity (77%; 95% CI, 70%-83%) and high specificity (93%; 95% CI, 88%-97%). Data sharing detection remained challenging; for instance, rtransparent detects data sharing with a sensitivity of 74% (95% CI, 68%-80%) and a specificity of 59% (95% CI, 46%-70%). Different diagnostic accuracies were observed depending on the type of research and the journal, likely due to different formatting standards. All results are shown in Table 25-1025. Limitations include the declarative nature of some practices (eg, data sharing).
Conclusions
Our study provides a detailed description of core open science practices across leading biomedical journals. It also highlights current challenges regarding the accuracy of automated tools in detecting these practices. While these tools likely provide valuable insights into overall practices, it is crucial to remain aware of the potential ranking biases introduced by these tools, as well as their limitations in providing detailed feedback for individual studies.
1Univ Rennes, Inserm, EHESP, Irset (Institut de recherche en santé, environnement et travail), UMR_S 1085, Rennes, France, constant.vinatier1@gmail.com; 2Department of Epidemiology and Data Science, Amsterdam University Medical Centers, Amsterdam, the Netherlands; 3QUEST Center for Responsible Research, Berlin Institute of Health at Charité–Universitätsmedizin Berlin, Berlin, Germany; 4Department of Epidemiology, Faculty of Health, Medicine, and Life Sciences, Maastricht University, Maastricht, the Netherlands; 5Bennett Institute for Applied Data Science, Nuffield Department of Primary Care Health Sciences, University of Oxford, Oxford, UK; 6Univ Rennes, IRISA, Inria, CNRS, Rennes, France; 7Department of Otorhinolaryngology and Head and Neck Surgery, University Medical Center Utrecht, Utrecht, the Netherlands; 8Brain Center, University Medical Center Utrecht, Utrecht, the Netherlands; 9Univ Rennes, CHU Rennes, Inserm, EHESP, Irset (Institut de recherche en santé, environnement et travail), UMR_S 1085, Rennes, France; 10Institut Universitaire de France (IUF), France.
Conflict of Interest Disclosures
None reported.
Funding/Support
As part of the OSIRIS project, this work was supported by the European Union’s Horizon Europe Research and Innovation Program under grant agreement number 101094725. Constant Vinatier, Ayu Putu Madri Dewi, Gowri Gopalakrishna, Nicholas J. DeVito, Inge Stegeman, Mariska M. G. Leeflang, and F. Naudet are members of this project.
Role of the Funder/Sponsor
The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
