Comparing Observational Exposure-Phenotype Correlations With Large Language Model Predictions
Abstract
Chirag J. Patel,1 Arjun K. Manrai,1 Randall J. Ellis,1 John P. A. Ioannidis2
Background
Large language models (LLMs) are increasingly used to synthesize biomedical evidence, yet it is unknown whether their inferred exposure-phenotype relationships mirror observational findings.1,2
Objective
To quantify concordance of exposure-phenotype empirical associations using National Health and Nutrition Examination Survey (NHANES) data with predictions from an LLM (ChatGPT 4o [OpenAI]).
Methods
We computed 115,056 exposure-phenotype partial correlations among 560 chemical, dietary, and clinical exposures and 268 phenotypes in 80,000 US adults (1999-2018 NHANES).3 We then sampled 1500 pairs: 500 significant after Bonferroni correction, 500 significant only after false-discovery rate (FDR) correction, and 500 not significant—to fit within the LLM prompt window. An OpenAI-o1-aug24 model (temperature = 0) acting as a prompted “epidemiologist” was asked to predict the sign, magnitude, and P value of each pair and rated the evidence generated as high, trending, or borderline. Verifiability of LLM-cited references was assessed by estimating κ values and Spearman correlation.
Results
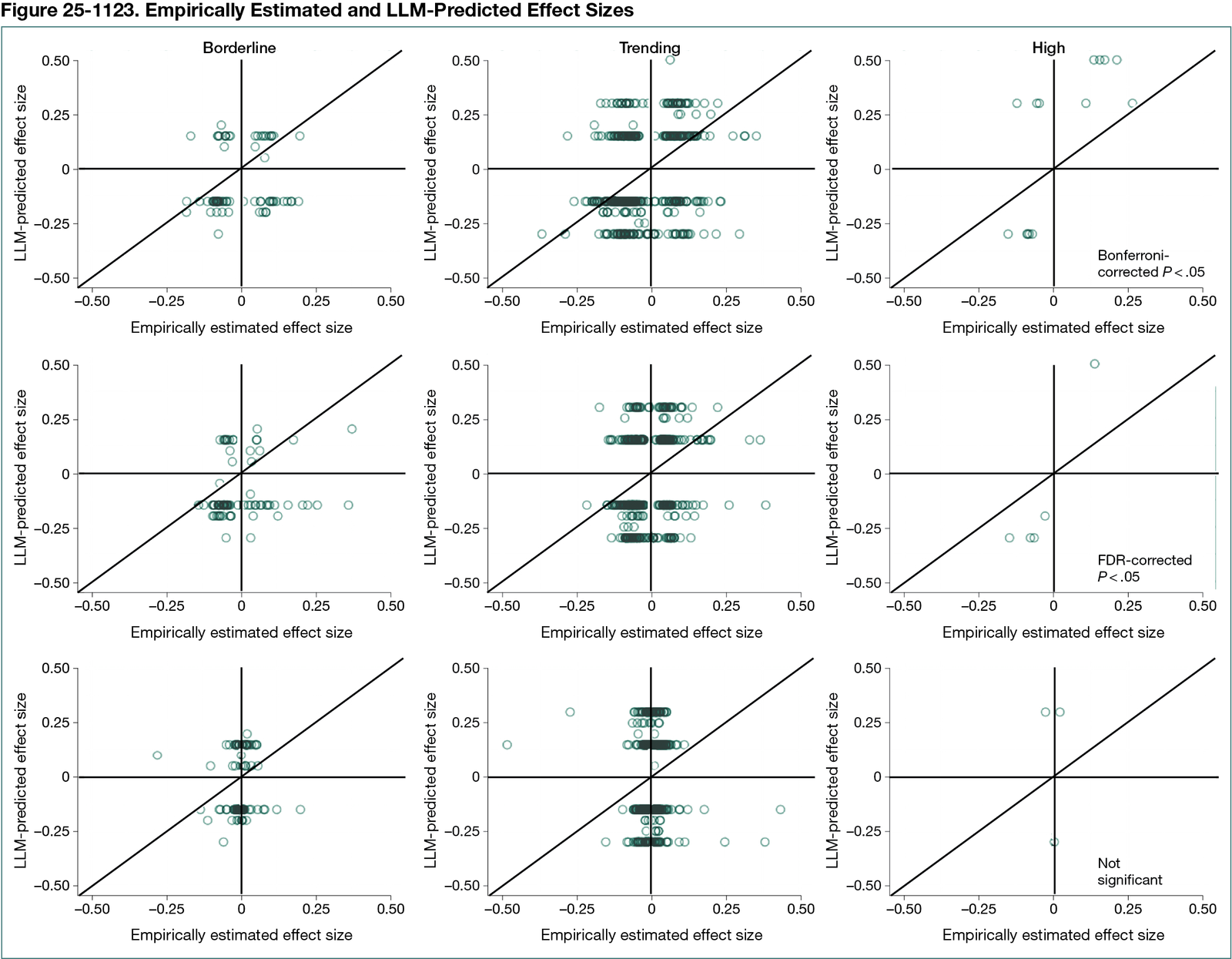
The LLM labeled 23 pairs as high evidence, 244 as trending evidence, and 1233 as borderline evidence. Sign concordance with NHANES was 73% for high-confidence pairs (κ = 0.49), 65% for Bonferroni pairs (κ = 0.25), 43% for FDR-only pairs (κ = 0.09), and 12% for borderline pairs (κ approximately 0) (Figure 25-1123). Random sign assignment yielded a κ of 0.0001. Concordance for correlation magnitudes was modest (Spearman ρ = 0.21). Permutation tests confirmed no greater-than-chance performance on nonsense data. Mechanistic explanations dominated high-confidence pairs; 41% of cited references (615 of 1500) were unverifiable.
Conclusions
LLMs captured directionality for well-established exposure-phenotype associations but faltered on borderline signals, highlighting both the promise and current limits of LLM-based evidence synthesis in exposomic epidemiology.
References
1. Ioannidis JPA, Loy EY, Poulton R, Chia KS. Researching genetic versus nongenetic determinants of disease: a comparison and proposed unification. Sci Transl Med. 2009;1(7):7ps8. doi:10.1126/scitranslmed.3000247
2. Patel CJ, Ioannidis JPA. Studying the elusive environment in large scale. JAMA. 2014;311(21):2173-2174. doi:10.1001/jama.2014.4129
3. Patel CJ, Rehkopf DH, Leppert JT, et al. Systematic evaluation of environmental and behavioural factors associated with all-cause mortality in the United States National Health and Nutrition Examination Survey. Int J Epidemiol. 2013;42(6):1795-1810. doi:10.1093/ije/dyt208
1Department of Biomedical Informatics, Harvard Medical School, Boston, MA, US, chirag@hms.harvard.edu; 2Department of Medicine, Epidemiology and Population Health, Stanford University School of Medicine, Stanford, CA, US.
Conflict of Interest Disclosures
John P. A. Ioannidis is a member of the Peer Review Congress Advisory Board but was not involved in the review or decision for this abstract.
Funding/Support
This study was funded by grants R01ES032470 and U24ES036819 from the National Institutes of Environmental Health Sciences and grant R01DK137993 from the National Institute of Diabetes and Digestive and Kidney Diseases.
Role of Funder/Sponsor
The sponsors had no role in this study.
